#30 Altered Cell Membranes, Ill-Equipped Ningaloo Coral and Snake Venom Evolution...
Scramblase rearranging cell membranes, ill equipped coral reefs in western Australia and the evolution of snake venom genes.
⭕Altered Membranes
How scramblase rearranges cell membranes…
The cell membrane is simply a biological membrane that separates the interior of all cells from the outside environment. A class of proteins known as TMEM16 scramblases permit rearrangement of lipids in the cell membrane by thinning the membrane, according to new research.
The model created by Weill Cornell Medicine investigators is based on high resolution images of the protein scramblase and challenged the prevailing theory of how the proteins carry out their fundamental role.
Scramblases are proteins responsible for the translocation of phospholipoids between the two monolayers of a lipid bilayer. Scramblase proteins reside in the outer membranes of almost all eukaryotic cells and can “scramble” the normal arrangement of fat related lipid molecules, a well known constituent of the membranes. Dysfunctions and disruption of the membrane by the TMEM16 scramblases is key in numerous biological events such as blood coagulation and membrane repair.
Cryogenic electron microscopy is a method by which samples are cooled to cryogenic temperatures. It allows macromolecular structure determination without the need for crystallization. Through using this method the team were able to analyse the TMEM16 scramblase at near-atomic scale resolution in a membrane like environment. The images suggested how the scramblases work…
“We’re proposing that the TMEM16 scramblase protein works not by interacting directly with specific lipids, but instead by mechanically altering the structure of the membrane, making it generally very thin and disordered,” said Alessio Accardi, professor of physiology and biophysics in anaesthesiology at Weill Cornell Medicine.
The outer membrane of the cell is composed of two tightly ordered layers of lipid molecules, with the outer and inner layers composes of distinct sets of lipid molecules types. A TMEM16 scramblase spans these layers and has a tube structure that is opened with the binding of calcium ions.
Activation of the protein allows lipids to lose a typical arrangement and distribute between the two layers. The previous and dominant understanding was that the chute like structure acted like the groove of a card reader with lipids moving freely between the two membrane leaflets to lose their usual ordering. However, the precise mechanism of lipid scrambling was unclear.
Through building a 3D model of this mechanism from a fungal species, it was found that the version of scramblase was similar to to that of human proteins. The imaging data implied that the ‘card reader’ model was incorrect. Rather than specific interactions with lipids, the scramblase chute locally deforms the membrane to make it significantly thinner, disorganizing lipids in the vicinity.
“We’re proposing that TMEM16 scramblase mechanically alters the structure of the membrane, making it very thin and disordered near the groove,” Accardi said. “This enables lipid movement without necessitating interactions with the inside of the groove, as in the credit card reader model.”
The team are now working to confirm their findings in mammalian versions of the scramblase and study the functions in regulated cells with an aim of developing small molecule drugs to restore function and treat disease.
🪸Ill-Equipped Coral?
Ningaloo corals ill-equipped for climate change…
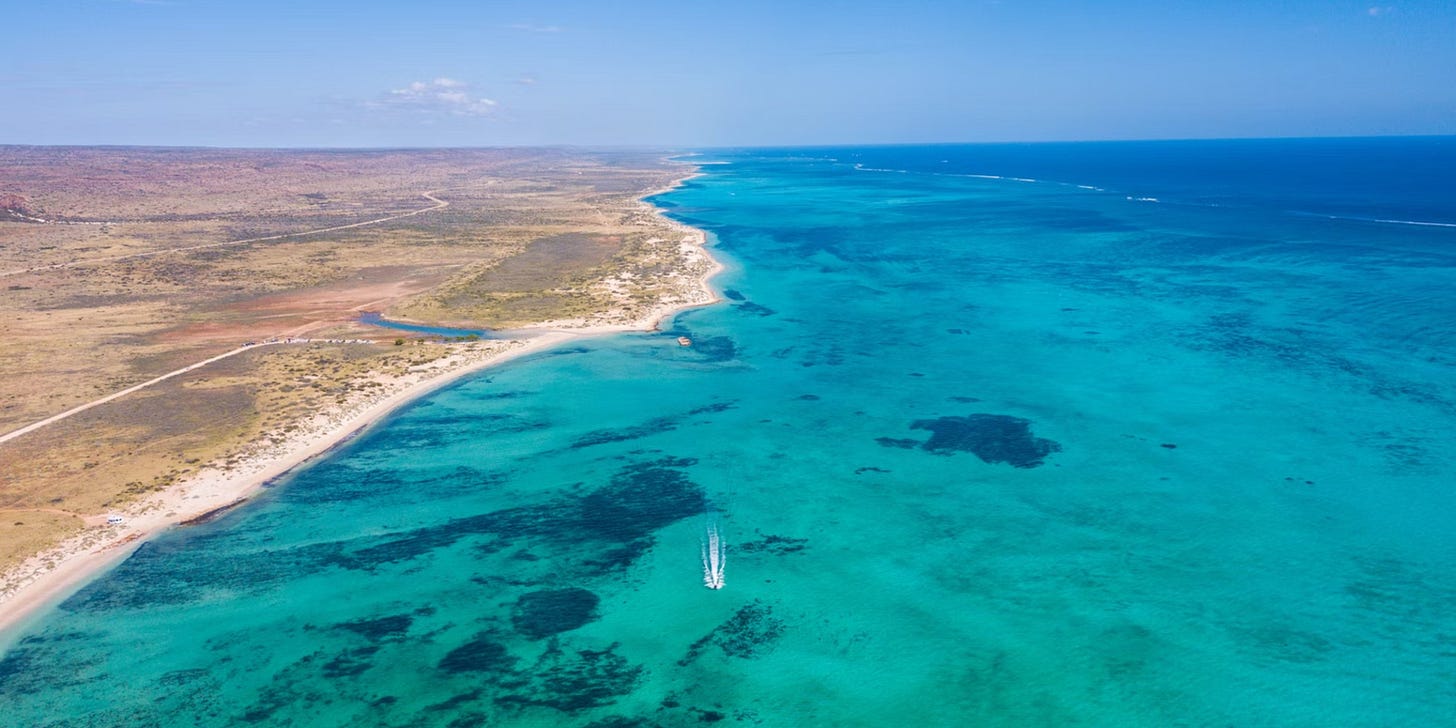
The west coast of Australia (WA) may differ from the prediction that all coral species would move towards cooler water to protect themselves.
The new study focused around investigating coral population connectivity and their adaptive capacity and found that corals growing in different reef systems are generally isolated from each other. The research team based their findings on genetic data of the reef building coral Acropora digitifera, from 5 different reef sites. The study aimed to uncover how connected these reef systems are, as well as their resiliency in relation to future climate scenarios.
Lead research PhD student Arne Adam, said how climate change had caused widespread loss of species biodiversity and ecosystem productivity across the globe, particularly on tropical coral reefs. He said the results suggest corals from northern reefs in WA are isolated from each other, meaning that corals may not be able to move to more southern reef regions. "Having segregated reefs means that it's hard for the corals to move between the regions. If corals at one reef die out, it is unlikely that this reef will be rescued by newcomer corals from neighbouring reefs," Mr Adam said.
He went on to say how previous research had indicated that the southern regions would become coral hotspots in the future, however, based on this data it is unknown that corals at southern locations have the needed genetic adaptations to survive the rapidly heating ocean.
"We found corals growing in northern reef regions such as the inshore Kimberley, including Adele Island and Beagle Reef, are better adapted to handle future ocean warming, whilst the coral community at Ningaloo Reef is in danger of losing diversity because they are not well-equipped to survive a warming ocean," said Mr Adam.
Dr Zoe Richards went on to say that for the Ningaloo reef system, this combination of traits could spell disaster under extreme future climate scenarios. This study helps to predict exactly which coral communities may be resilient or vulnerable to future climate change and allows us to adapt and react accordingly.
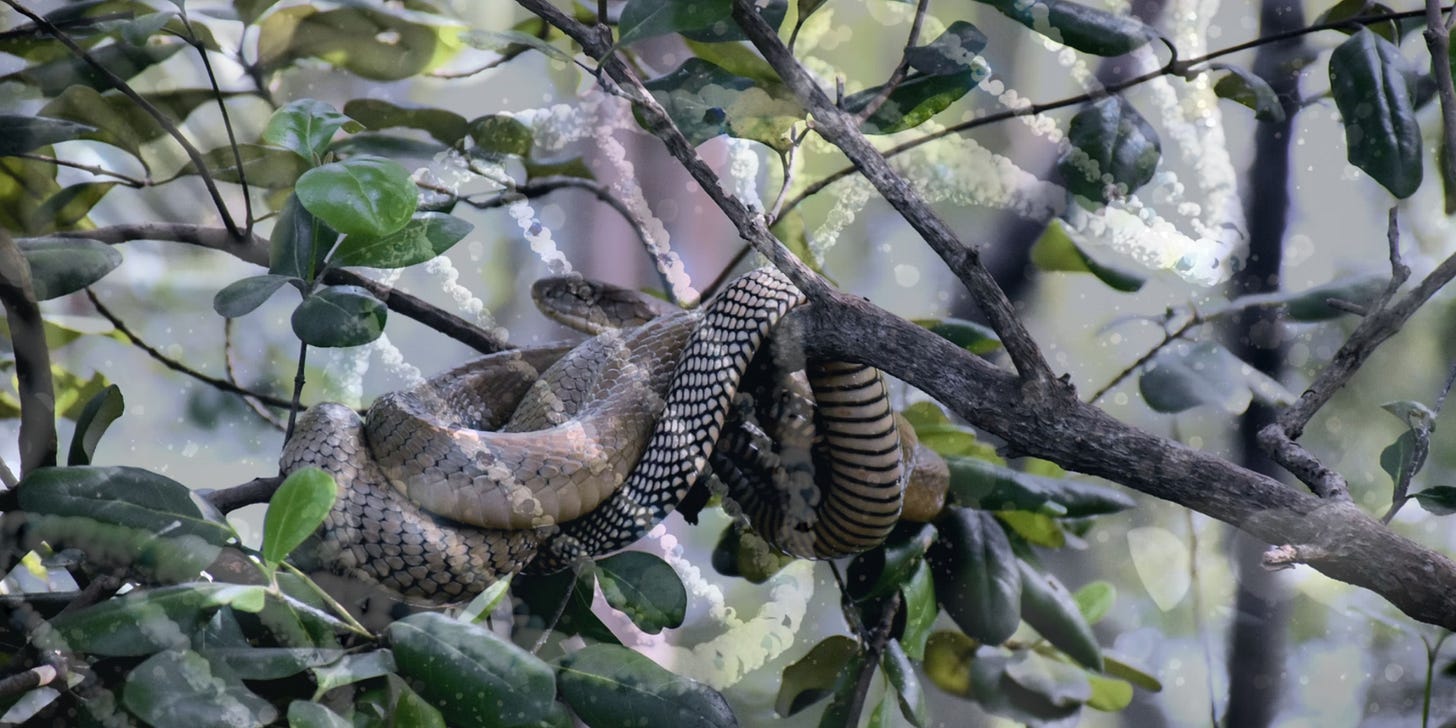
🐍Snake Venom Evolution
Breakthrough into snake venom gene evolution…
A recent study from the University of Texas at Arlington provides the first comprehensive explanation of how snake venom regulatory systems evolved, shining a light on how the evolution of new complex traits arises.
“We have relatively few detailed examples of how new regulatory systems evolve to drive novel complex traits,” Todd Castoe, author of the paper, said. “This study provides a valuable example that illustrates a surprising number of distinct ‘strategies’ for how evolution may rewire regulatory networks, providing key expectations for how such rewiring may occur in other species, including humans.”
Ever since the first proposal of natural selection, the scientific community has not been able to identify exactly how complex traits, such as snake venom, evolve.
“This work gives us a better understanding of how snake venom evolved and how venom production functions at a genomic level,” said Blair Perry, a UTA alumnus and postdoctoral researcher in the School of Biological Sciences at Washington State University and lead author of the new paper. “In addition to studying specific venom genes, we can now investigate parts of the genome involved in the regulation of these genes as well,” said Perry. “This opens up new opportunities to understand how variation in snake venom, both within and between snake species, corresponds to variation in the genome.”
2019 saw the World Health Organisation declare snakebites as a neglected tropical disease with the main challenge in terms of treatment being their extreme variation. Evolution in snake venom required snakes to develop highly specialized venom glands to produce and store a diverse and deadly protein concoction. Venom glands are thought to have evolved from ancestral salivary glands, but Perry and colleagues show that this also required the evolution of new regulatory sequences and the repurposing of existing regulatory systems.
The teams earlier work had demonstrated that several mechanisms exist that promote this long-term storage, but the processes leading to the regulation, evolution and diversification of these systems have largely remained unknown. This new study demonstrates that in addition to the toxin genes, regulatory pathways common to vertebrate animals were also co-opted to control this system.
In this study the authors apply a diverse array of cutting-edge approaches to investigate this topic, and their ensuing collection of regulatory sequence, transcription factor, signalling cascade and chromatin accessibility data and associated analyses provides completely unparalleled insight into the regulatory architecture of the snake venom system,” said Nicholas Casewell an expert in the field, not associated with the research. “Their findings represent a major step forward in helping us to better understand how genes associated with internal physiological processes can be repurposed for external use in the form of venom.”
“By combining multiple types of genomic data, we were able to gain a more complete understanding of the diverse factors that play a role in the regulation and evolution of venom genes,” Perry said. “More broadly, this work provides a valuable example of the idiosyncrasies of evolution. One might expect that all venom genes followed the same evolutionary ‘strategies’ to become involved in venom. Our findings instead suggest that remarkably different genomic and evolutionary processes played critical roles in the evolution of specific venom genes.”
Weekly Topics
🏞️ Environmental
Galapagos giant tortoise is not actually extinct
Harvested - critically endangered fruit
Homogenisation threat to forest ecosystems
🦭Marine
How to avoid great white sharks and shark research
Why do female octopuses self destruct?
🐼 Conservation
Ugly reef fishes most in need of conservation
Red panda being driven to extinction
World/s most endangered sea turtle nests…
🦠 Disease and Illness
Host gut resistome in Gulf war
A map of our cells could help treat diseases
Mechanotransduction: Using nuclear mechanics to understand health
😷 COVID
Underlying biology of long COVID
West Africa cases underreported…
Machine learning tool can predict COVID variants
🧪 Biochemistry
Paramagnetic encoding of molecules
Anti-cancer mitochondrial drug found to be effective against most carcinoma cell lines
Decoding a key part of the cell…
🔬 Evolution
New insights on link between genetic mutations and biological evolution
How electric fish were able to evolve electric organs…
Recurring brain tumours shaped by genetic evolution and microenvironment
🧬 Genetics
Silent genetic mutations are harmful not neutral?
CRISPR based map ties every human gene to its function
Study sheds light on the molecular biology behind genetic epilepsy
📷 Weekly Camera Roll
Click on the text below to keep reading…
Reference List
Content may be adapted and edited for style and length.
⭕Altered Membranes
Falzone, M., Feng, Z., Alvarenga, O., Pan, Y., Lee, B., Cheng, X., Fortea, E., Scheuring, S. and Accardi, A., 2022. TMEM16 scramblases thin the membrane to enable lipid scrambling. Nature Communications, 13(1).
News Release. https://news.cornell.edu/stories/2022/06/study-sheds-light-how-scramblase-proteins-rearrange-cell-membranes
🪸Ill-Equipped Coral?
Adam, A., Thomas, L., Underwood, J., Gilmour, J. and Richards, Z., 2022. Population connectivity and genetic offset in the spawning coral <i>Acropora digitifera</i> in Western Australia. Molecular Ecology,.
News Release. https://research.curtin.edu.au/news/ningaloo-corals-are-ill-equipped-to-handle-future-climate-change/?type=media
🐍Snake Venom Evolution
Perry, B., Gopalan, S., Pasquesi, G., Schield, D., Westfall, A., Smith, C., Koludarov, I., Chippindale, P., Pellegrino, M., Chuong, E., Mackessy, S. and Castoe, T., 2022. Snake venom gene expression is coordinated by novel regulatory architecture and the integration of multiple co-opted vertebrate pathways. Genome Research,.
News Release - https://www.uta.edu/news/news-releases/2022/06/01/breakthrough-study-examines-evolution-of-snake-venom-genes